It provides a ggplot of the sequence ratios of index and marker cohorts.
Arguments
- result
Table output from summariseSequenceRatios.
- onlyASR
If set to be TRUE then only adjusted SR will be plotted. Otherwise if it is set to be FALSE then both adjusted and crude SR will be plotted.
- plotTitle
Title of the plot, if NULL no title will be included in the plot.
- labs
Axis labels for the plot.
- colours
Colours for sequence ratio.
- facet
The variable to facet by.
Examples
# \donttest{
library(CohortSymmetry)
cdm <- mockCohortSymmetry()
cdm <- generateSequenceCohortSet(cdm = cdm,
indexTable = "cohort_1",
markerTable = "cohort_2",
name = "joined_cohort")
sequence_ratio <- summariseSequenceRatios(cohort = cdm$joined_cohort)
#> Warning: For at least some combinations, index is always before marker or marker always
#> before index
#> -- 5 combinations of 8 had index always before marker
#> -- 5 combinations of 8 had marker always before index
plotSequenceRatios(result = sequence_ratio)
#> Warning: Removed 3 rows containing missing values or values outside the scale range
#> (`geom_point()`).
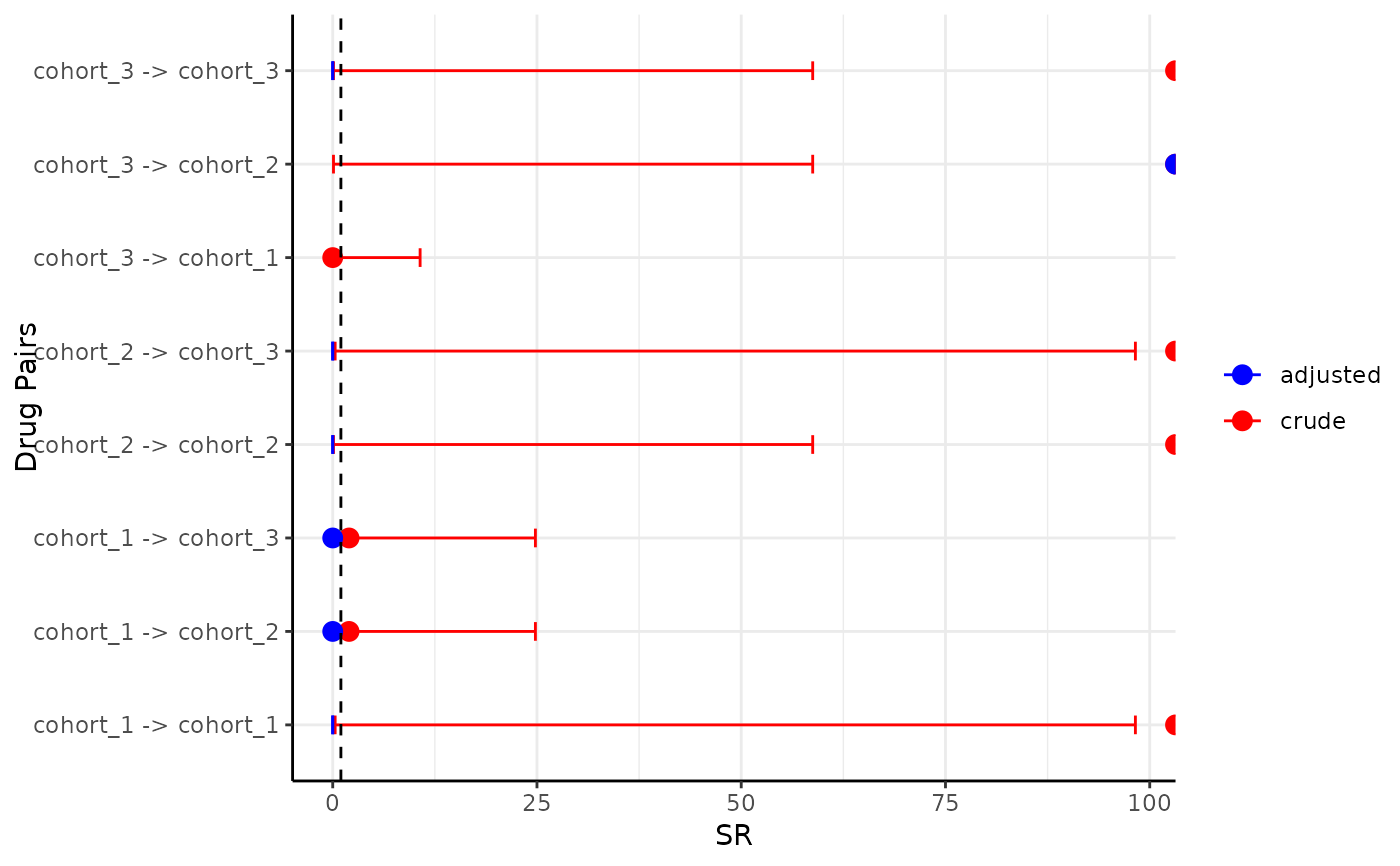 CDMConnector::cdmDisconnect(cdm = cdm)
# }
CDMConnector::cdmDisconnect(cdm = cdm)
# }
